Our lab develops synthetic biology and genomics approaches to map and engineer cell fate. We are interested in early embryogenesis, neurodevelopment, and cancer as models of cell fate. To study these systems, we use molecular engineering, genome engineering, computational modeling, as well as spatial and single-cell sequencing approaches.
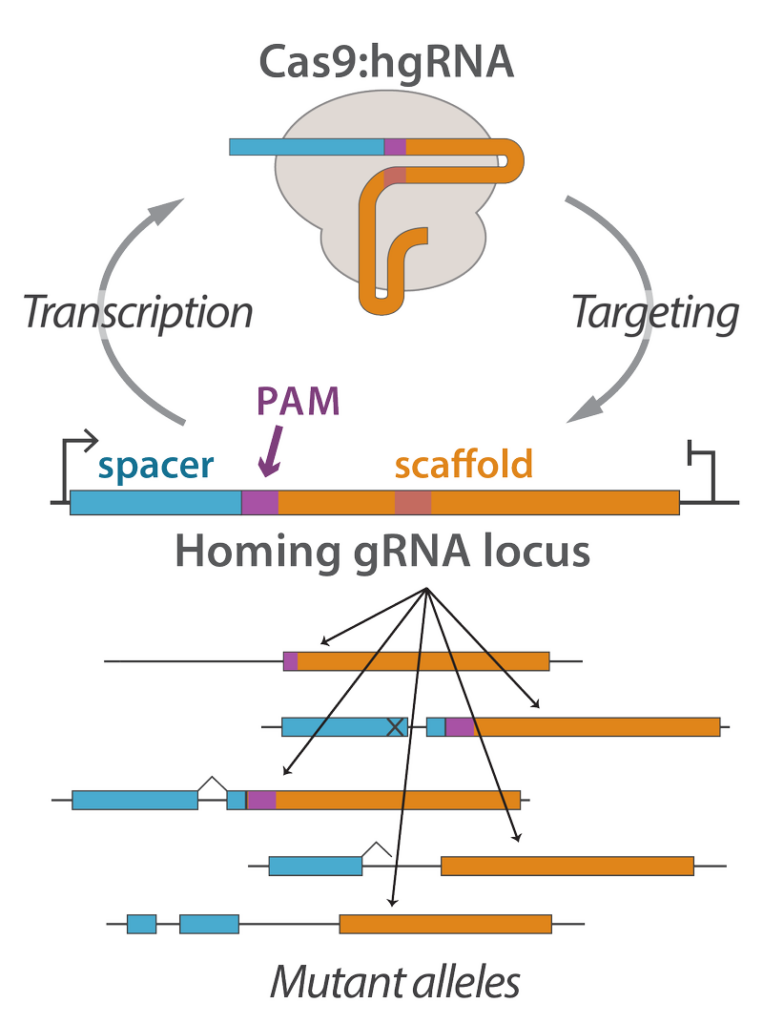
Molecular devices for genomic recording
Engineering cells to label their genomes with unique barcodes is an effective way of tracking cellular identity, behavior, and history. We are interested in developing molecular devices that accomplish this in vivo barcoding with both high efficiency and minimal negative side-effects. For instance, we have developed the homing CRISPR system as a versatile platform for in vivo barcoding. In this platform, homing guide RNAs (hgRNAs) target their own loci to create highly diverse and unique random mutations. Among other topics, we are applying these molecular devices in brain connectome mapping and the deep lineage tracing of embryonic development.
Further reading: Rapidly evolving homing CRISPR barcodes.
Cell fate mapping and lineage barcoding
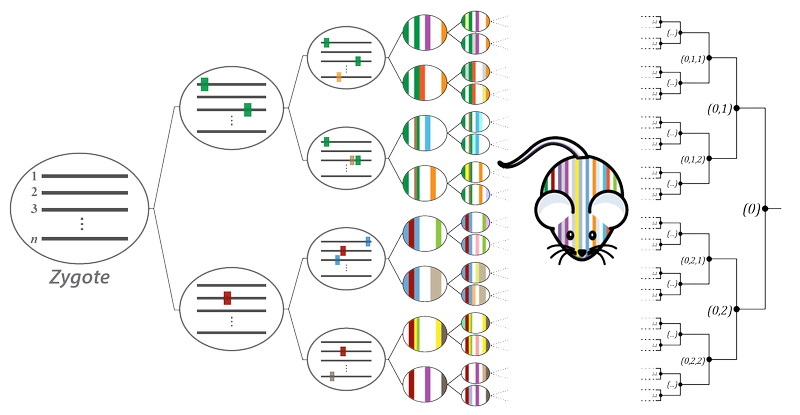
The development of an organism from single totipotent zygote is a subject of foundational interest for our lab. The path of orchestrated zygotic divisions and differentiation events during development creates cell lineages, much like a tree. This tree has been entirely mapped in C. elegans, an organism of ~1000 cells, and led to broad-ranging genetic insights. Our work aims to create a similar mapping system in higher eukaryotes, enabling characterization of deep, single-cell resolution lineage trees in mammals using molecular recording devices.
To this end, we have established a unique mouse model, dubbed MARC1 (Mouse for Actively Recording Cells!), which carries multiple hgRNA loci distributed across its genome. Activating these hgRNAs in the zygote results in their mutagenesis throughout gestation, creating developmentally barcoded mice in which lineage information is recorded in hgRNA barcodes. We use this strategy to study early and extraembryonic development, as well as the development of the central nervous system.
Further reading: Developmental barcoding of whole mouse via homing CRISPR.
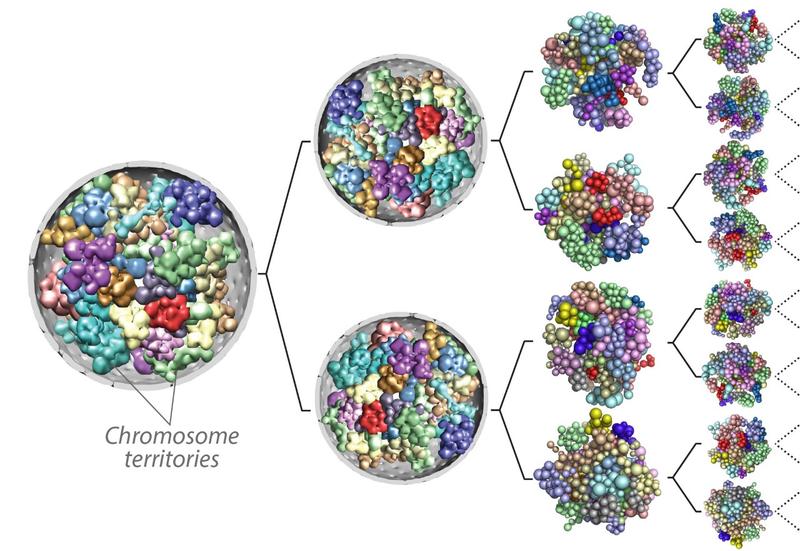
Genome 3D organization and cell fate specification
During embryogenesis, pluripotent cells undergo a succession of lineage commitments in a highly programmed fashion as they first differentiate into primary germ layers—the endoderm, mesoderm, and ectoderm—and then to all the cell types that compose the body. These differentiation events involve extensive reorganization of the epigenome and the genome three-dimensional (3D) structure. We are interested in the role of genome 3D structure in fate determination, the variability of that structure, and its interplay with the epigenome during embryogenesis. To this end, we are studying the 3D dynamics of genome reorganization during gastrulation and how these structures respond to disruptions in the epigenetic landscape. We are also examining how the genome recapitulates its 3D state after mitosis.
In situ readout of nucleic acids
The ability to extract transcriptomic, genomic, and epigenetic information from individual cells in high throughput while preserving detailed information about their spatial position enhances our understanding of biological behavior, especially in heterogenous mixtures such as tissues. We work on technologies that allow for high throughput readout of DNA and RNA sequences from fixed cells and tissue sections directly (in situ sequencing). We are particularly interested in using hybridization and sequencing-based readouts for in situ readout of nucleic acid barcodes in tissue sections.
Further reading: Deconvolving organogenesis in space and time via spatial transcriptomics in thick tissues.